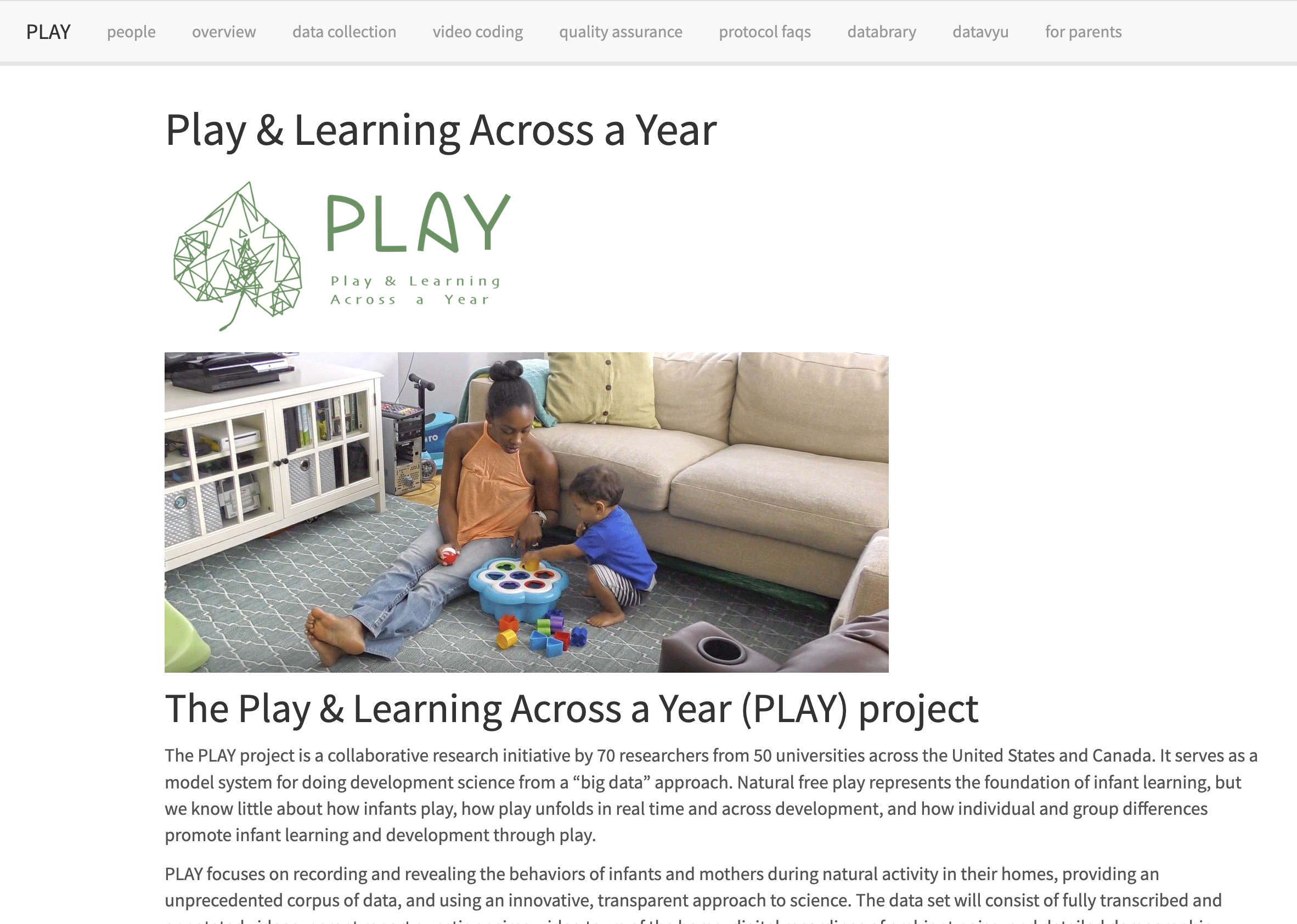
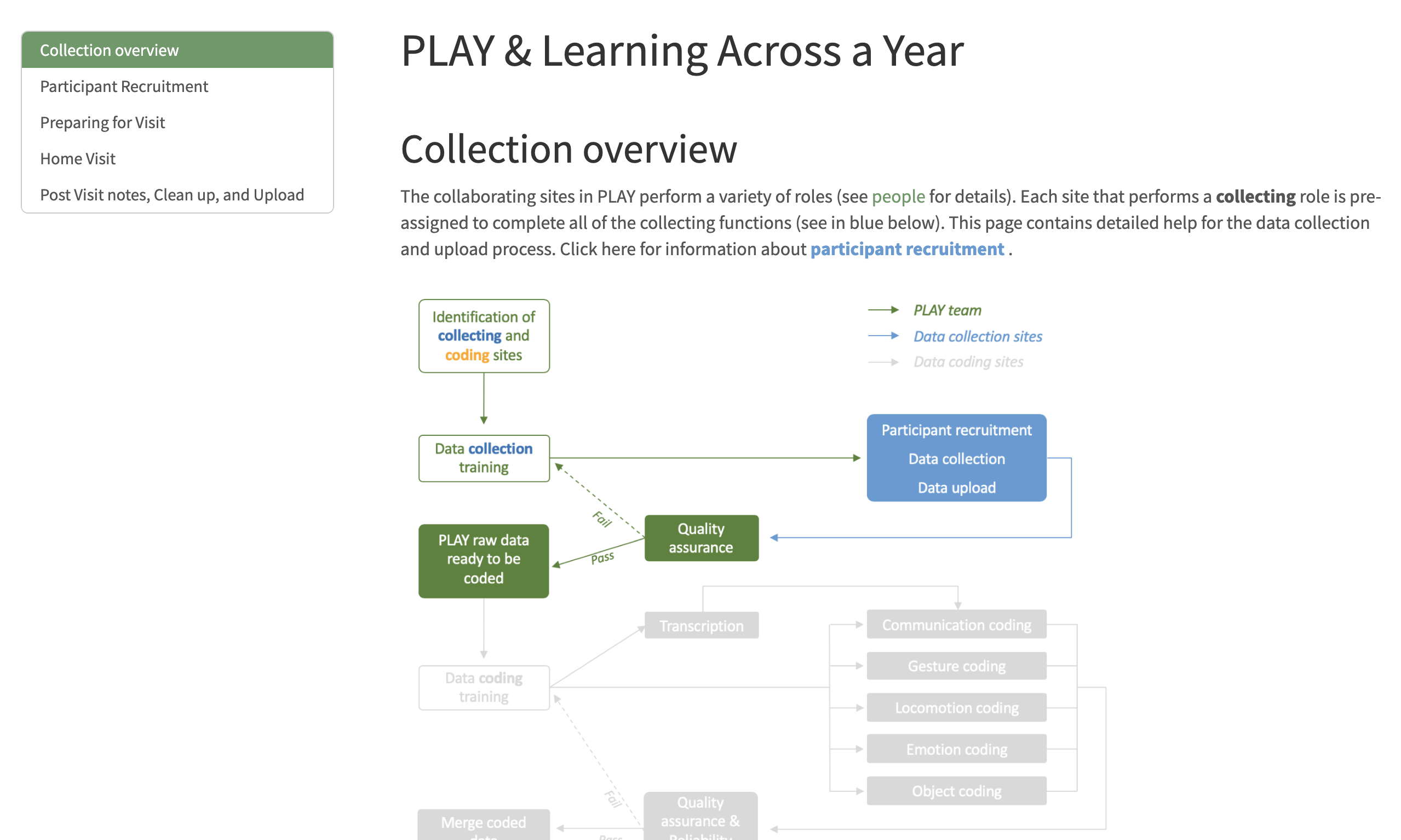
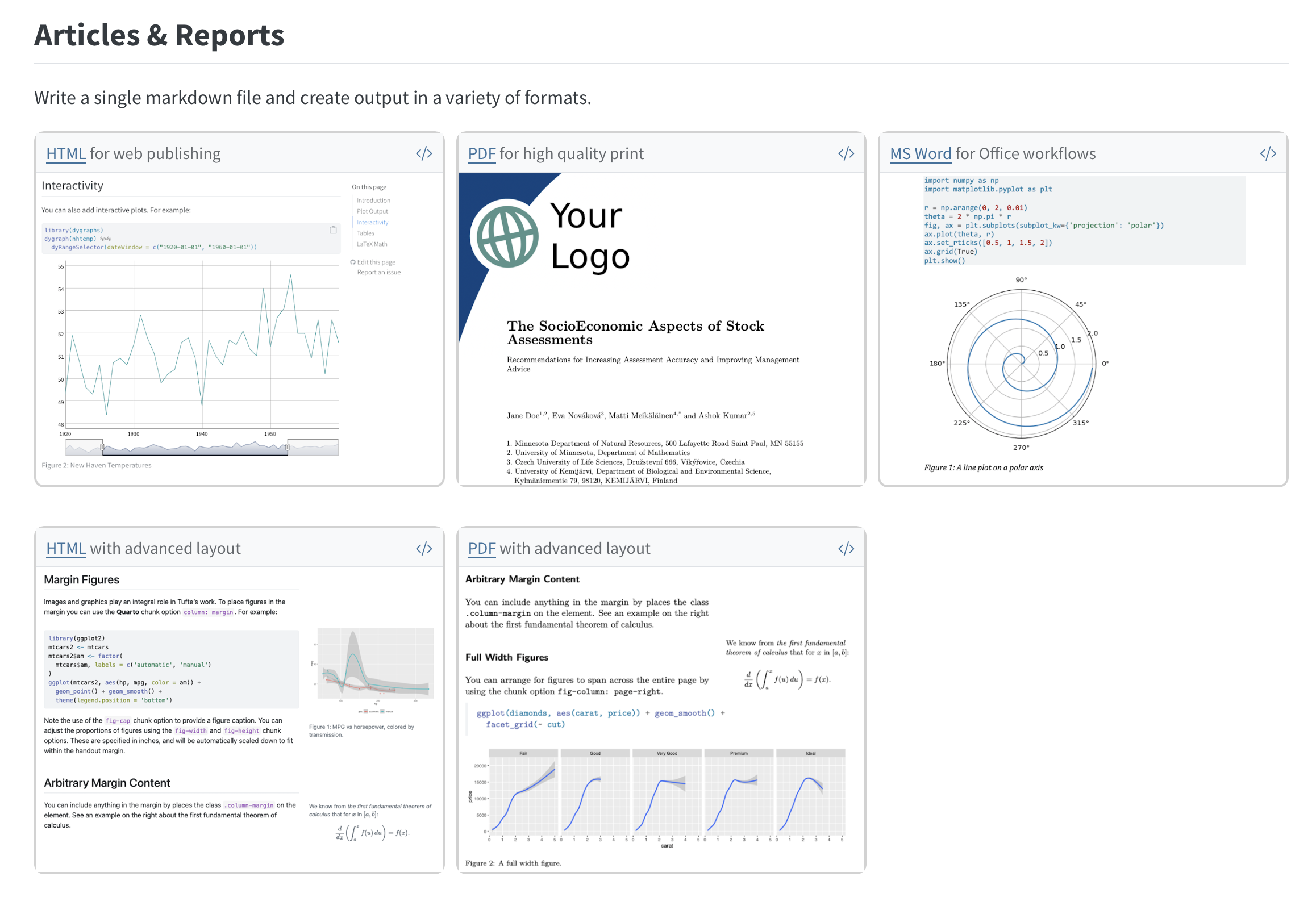
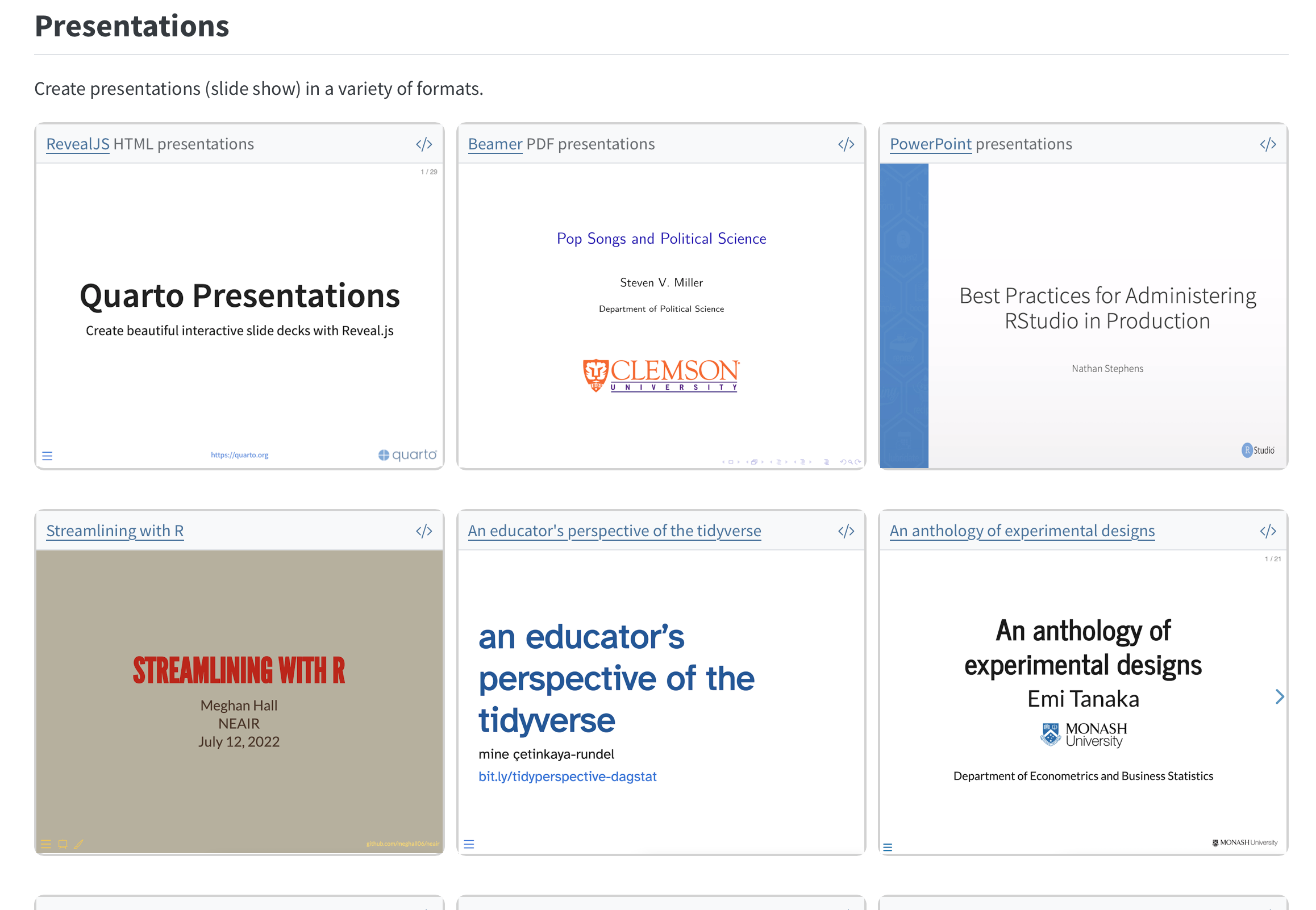
Quarto: Part II
Making useful things reproducibly
2026-05-10
Preliminaries
Follow-along
Agenda
- Why reproducibility?
- Why we should care
- Requirements for reproducibility
- What useful things?
- Case studies
- What’s next?
Why reproducibility
What R we talking about?
- Findings should be reproducible
- Same data + same code -> same results
- Findings should be robust
- Same data + new analysis -> comparable results & conclusions
- Findings should be replicable
- New data -> comparable results & conclusions
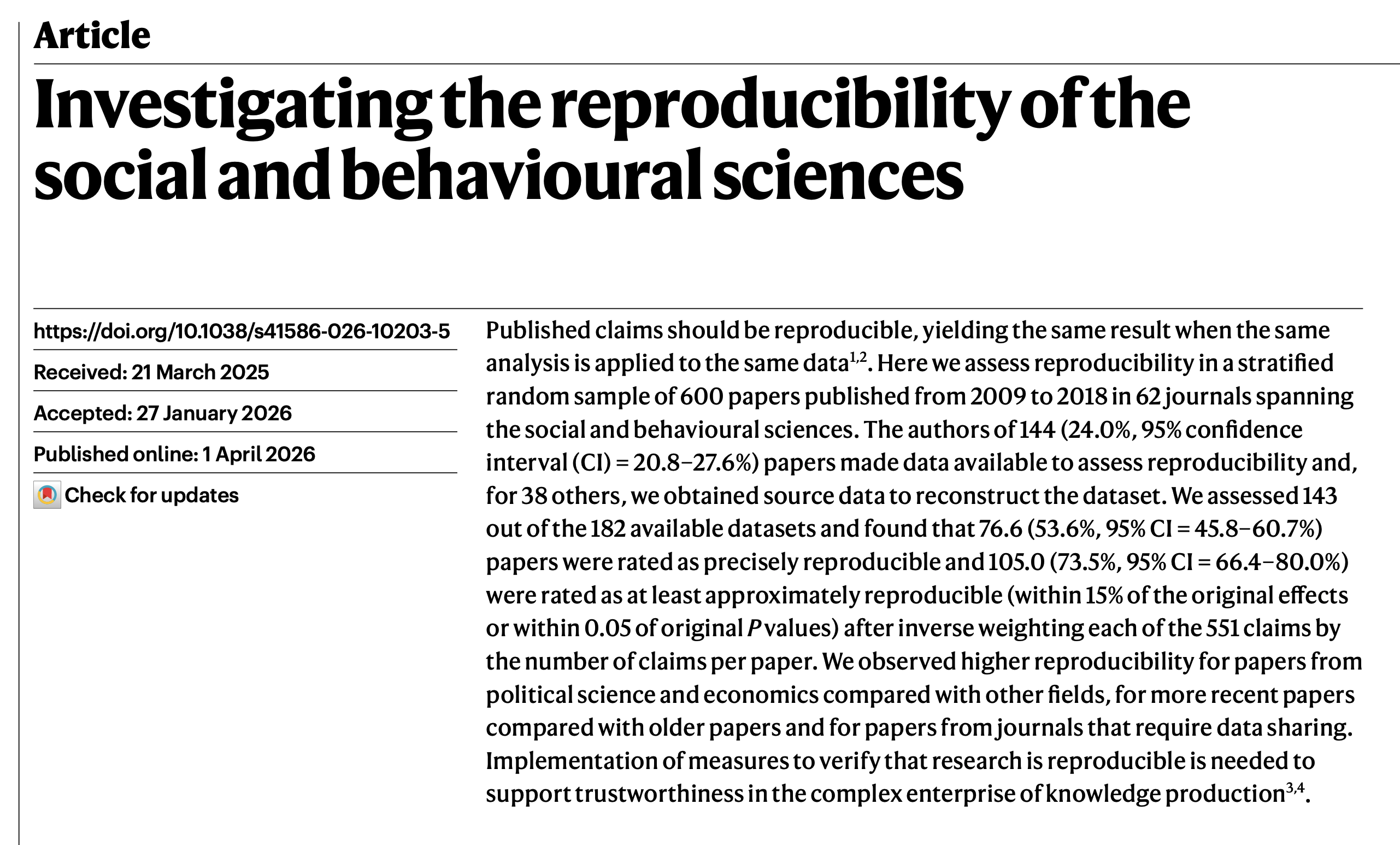
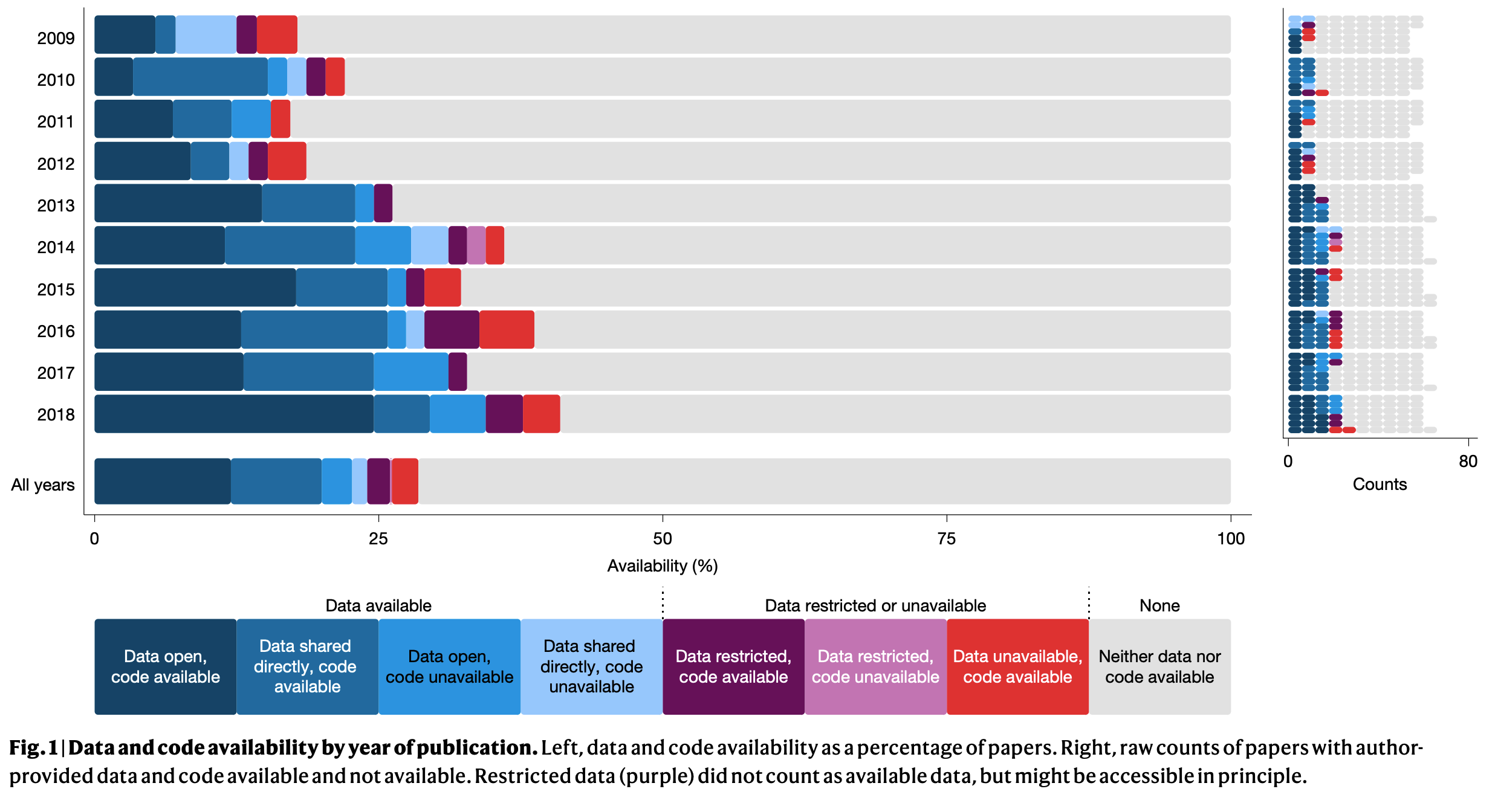
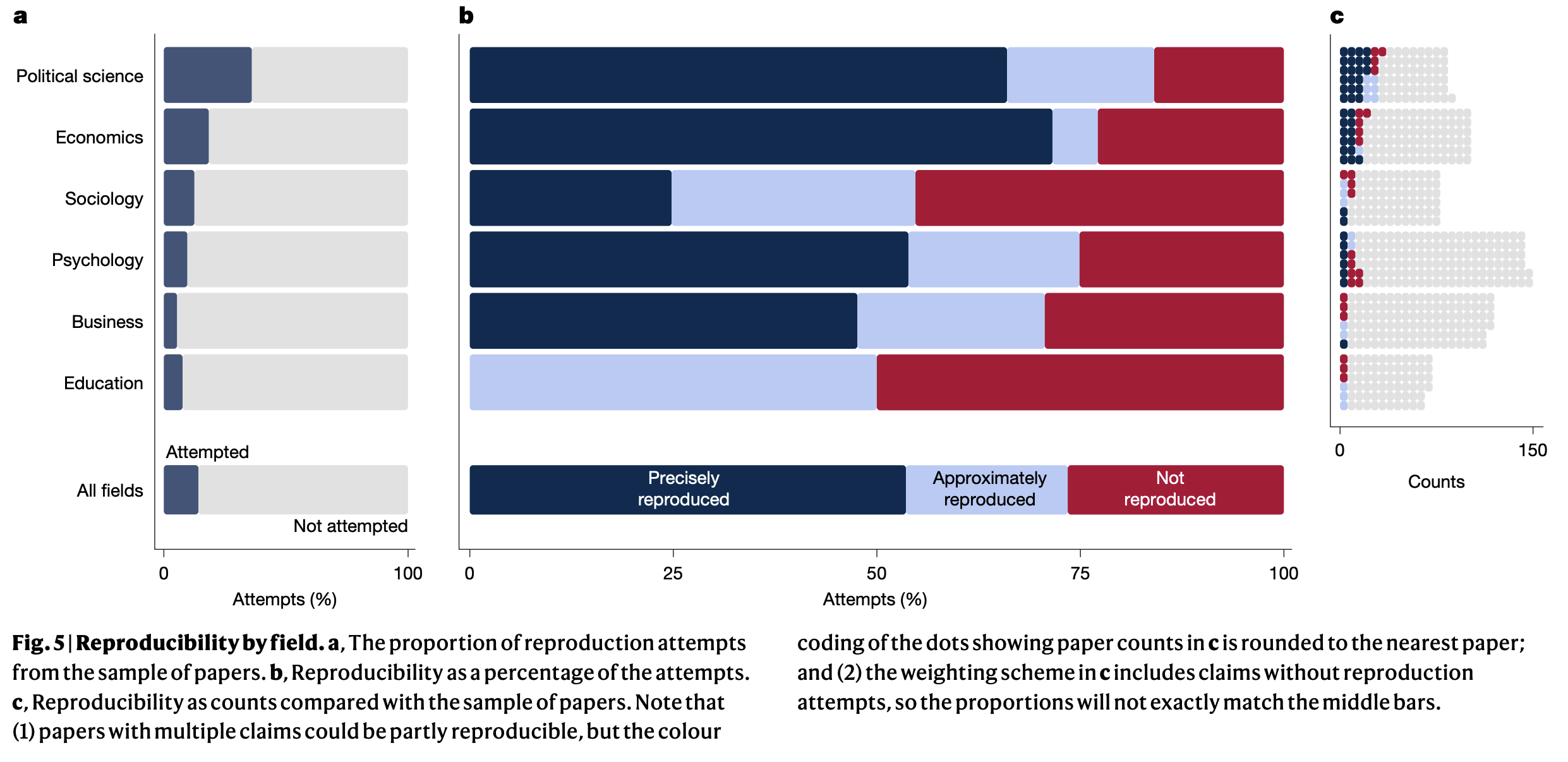
Miske et al. (2026)
We assessed 143 out of the 182 available datasets and found that 76.6 (53.6%, 95% CI=45.8–60.7%) papers were rated as precisely reproducible…
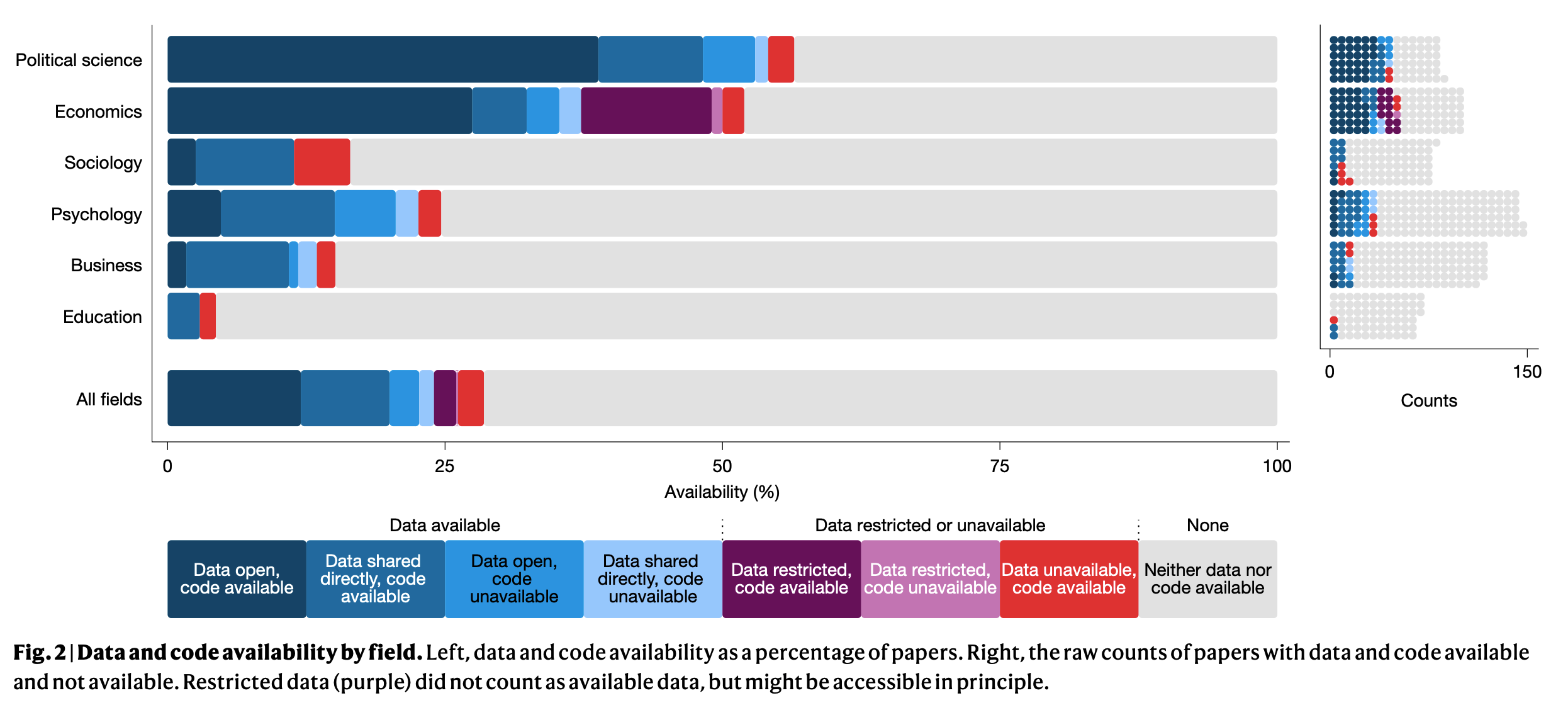
Miske et al. (2026)
…and 105.0 (73.5%, 95% CI=66.4–80.0%) were rated as at least approximately reproducible…
Miske et al. (2026)
…Implementation of measures to verify that research is reproducible is needed to support trustworthiness in the complex enterprise of knowledge production.
…The initial aim of the project was to repeat 193 experiments from 53 high-impact papers…However, the various barriers and challenges we encountered while designing and conducting the experiments meant that we were only able to repeat 50 experiments from 23 papers…
Errington, Denis, Perfito, Iorns, & Nosek (2021)
…the data needed to compute effect sizes and conduct power analyses was publicly accessible for just 4 of 193 experiments…none of the 193 experiments were described in sufficient detail in the original paper to enable us to design protocols to repeat the experiments…
Errington et al. (2021)
…While authors were extremely or very helpful for 41% of experiments, they were minimally helpful for 9% of experiments, and not at all helpful (or did not respond to us) for 32% of experiments…
Errington et al. (2021)
…This experience draws attention to a basic and fundamental concern about replication – it is hard to assess whether reported findings are credible.
Errington et al. (2021)
Why we should care
“The first principle is that you must not fool yourself—and you are the easiest person to fool. So you have to be very careful about that. After you’ve not fooled yourself, it’s easy not to fool other scientists.”
Feynman (1974)
Houses of straw, sticks, or
Stone?
What’s your “bus number”?
Building for reproducibility
Capture the workflow
- Humans
- Checklists, Standard Operating Procedures (SOPs), Protocols
- Quality Assurance (QA) checks
Requirements for reproducibility
- Humans
- Checklists, Standard Operating Procedures (SOPs), Protocols
- Quality Assurance (QA) checks
- Computers
- Data + code
- Package/library version management
- Testing (unit, regression, etc.)
- Consistent random number seeds, e.g.,
set.seed()
Principles for reproducibility
- DRY WIT
- Don’t Repeat Yourself
- Write It Down
Share your workflow
What useful things?
flowchart LR
A(["Idea"]) --> B["Proposal"]
B --> C("Project_1")
B --> D("Project_2")
C --> E["IRB_protocol"]
D --> E
C --> F["Lab_protocol"]
D --> F
C --> G["Data_pipeline_1"]
D --> H["Data_pipeline_2"]
B --> I["Conference_presentation"]
F --> I
G --> I
C --> I
I --> J["Journal_manuscript"]
H --> K["Lab_mtg_report"]
K --> J
F --> J
flowchart TD A["Grant proposal"] --> B["IRB Protocol"] B --> C["Lab Protocol"] C -->|"updates"| B C --> D["Data pipeline"] A --> E["Data management & sharing plan"] E --> B & C & D
flowchart TD A["Grant proposal"] --> B["IRB Protocol"] B --> C["Lab Protocol"] C --> B C --> D["Data pipeline"] A --> E["Data management & sharing plan"] E --> B & C & D A & E --> F["*.docx"] B --> F C --> F C --> G["*.pdf"] D --> nodeX:::hidden classDef hidden display:none
One framework to rule them all…
- Write text files
- Render to
- HTML
- Word (docx), Powerpoint (pptx)
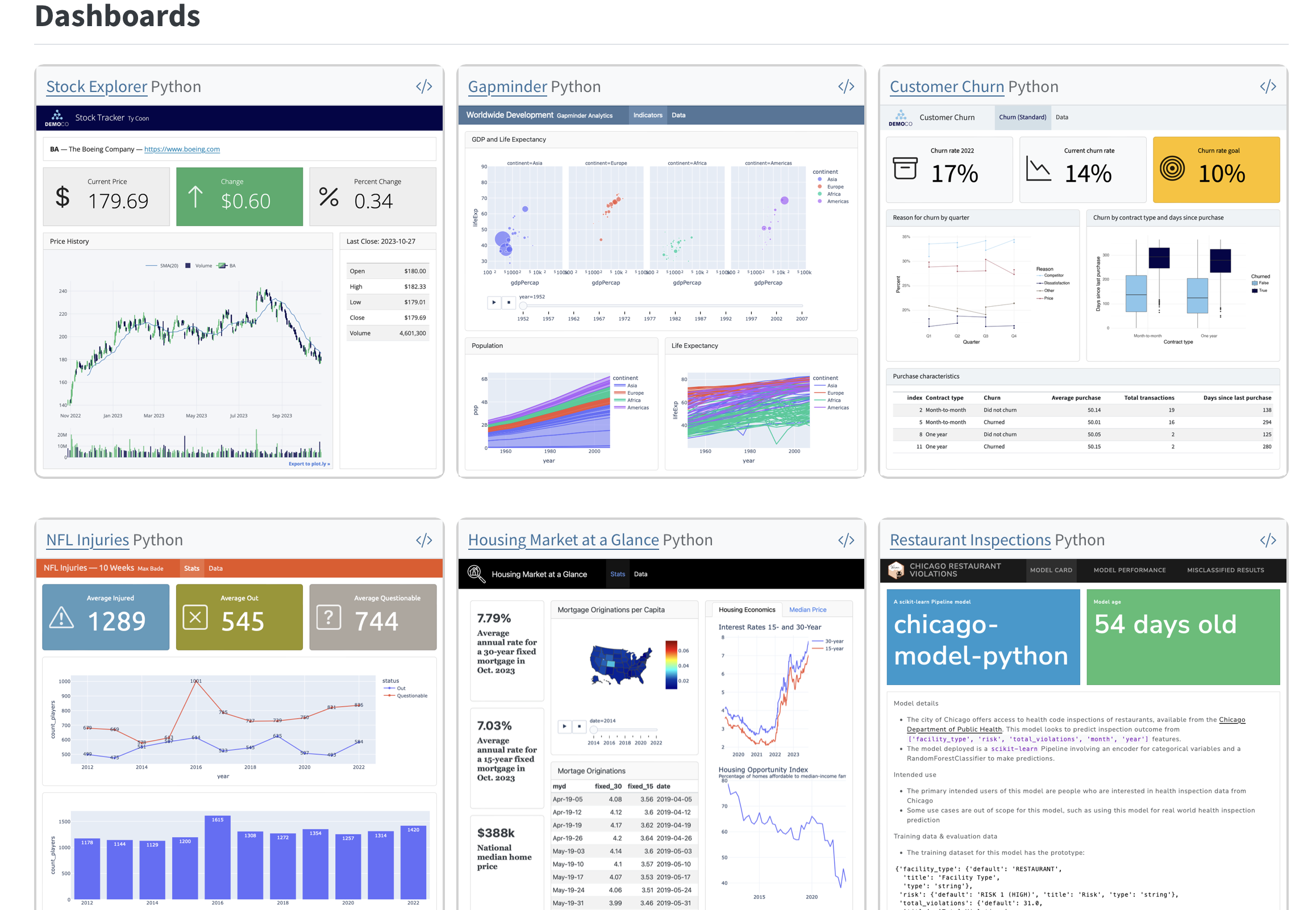
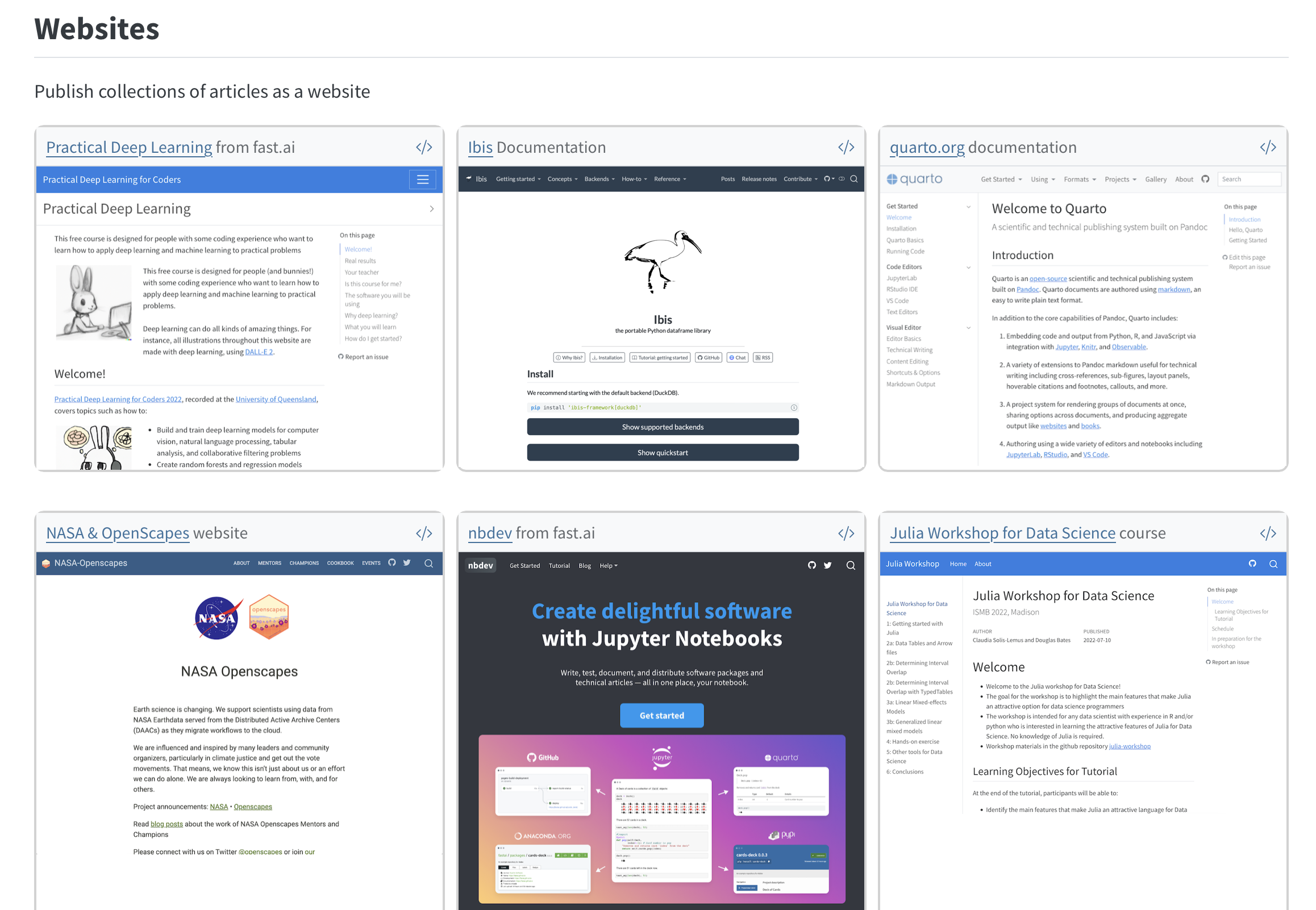
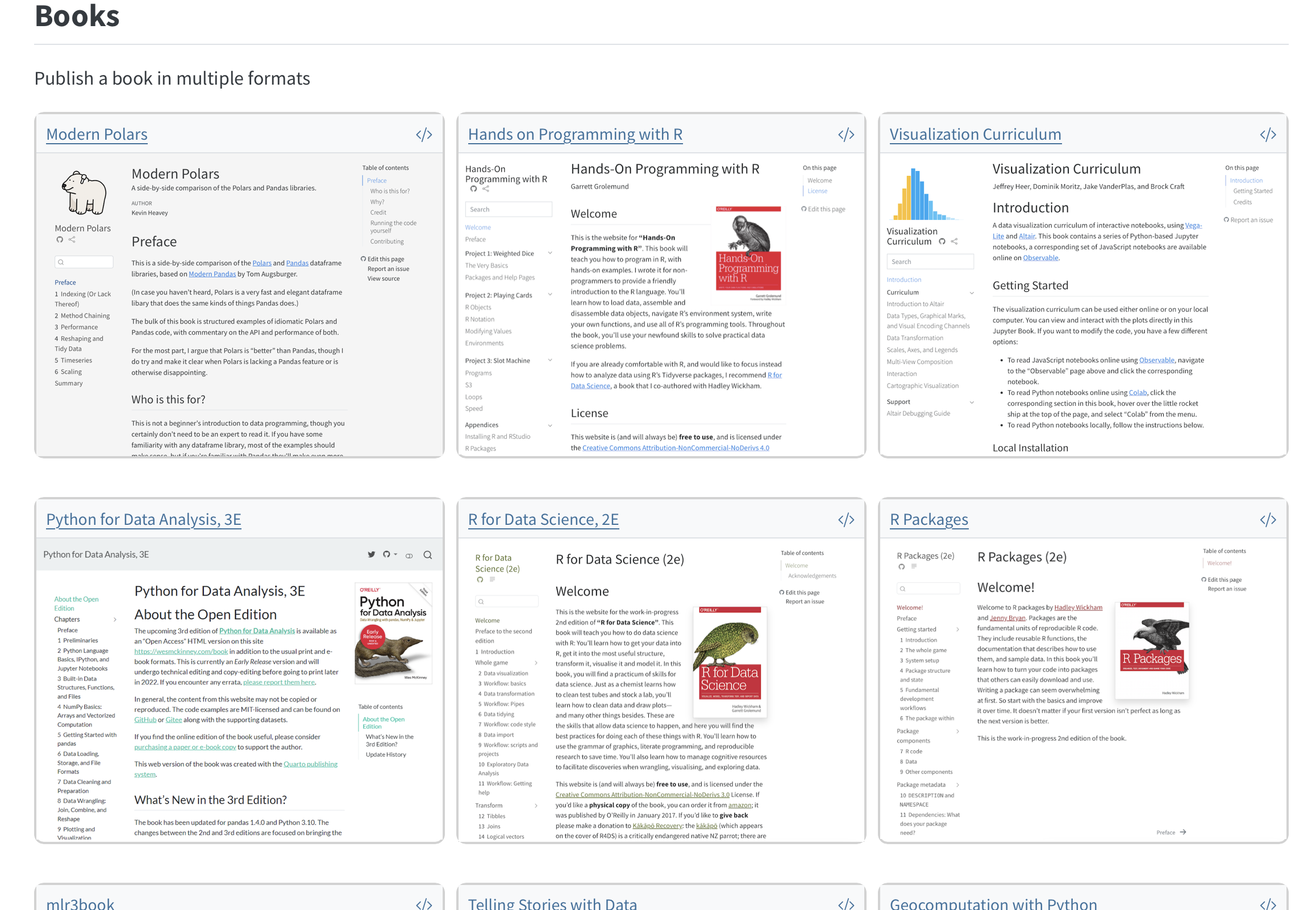
What Quarto supports
Things I’ve built
- Blogs
- Data cleaning, analysis, & visualization workflows
- Course/workshop sites, one-off talks
- Posters
- Interactive dashboards
Why HTML?
- Easily shared
- Cheaply shared1
- DRY WIT
- Meet emerging accessibility standards
- Machine-readable
- Might help other humans
Case studies
Data pipeline
- Gathering
- Cleaning
- Visualizing
- Analyzing
- Documenting/sharing
flowchart TD A["Gathering"] --> B["Cleaning"] B --> C["Visualizing"] C --> B C --> D["Analyzing"] D --> B
Quarto data pipeline
- One Quarto document (
*.qmd)- Separate sections for each step.
- Separate documents (Gather/Clean, Visualize).
Gathering: In general
- Automate!
- Application Programming Interfaces (APIs)
- How to talk to Qualtrics, Google Forms, RedCAP, etc.
- R packages: {qualtRics}, {googlesheets4}, {REDCapR}
- Leave source/raw data intact
- Clean “downstream”
- Automate cleaning
Gathering: In general
- Credentials
- Store securely
- Not with data analyses!
- Quarto projects: Locally in
~/.Renviron
- Where to save/store data
- Individual session/person or group
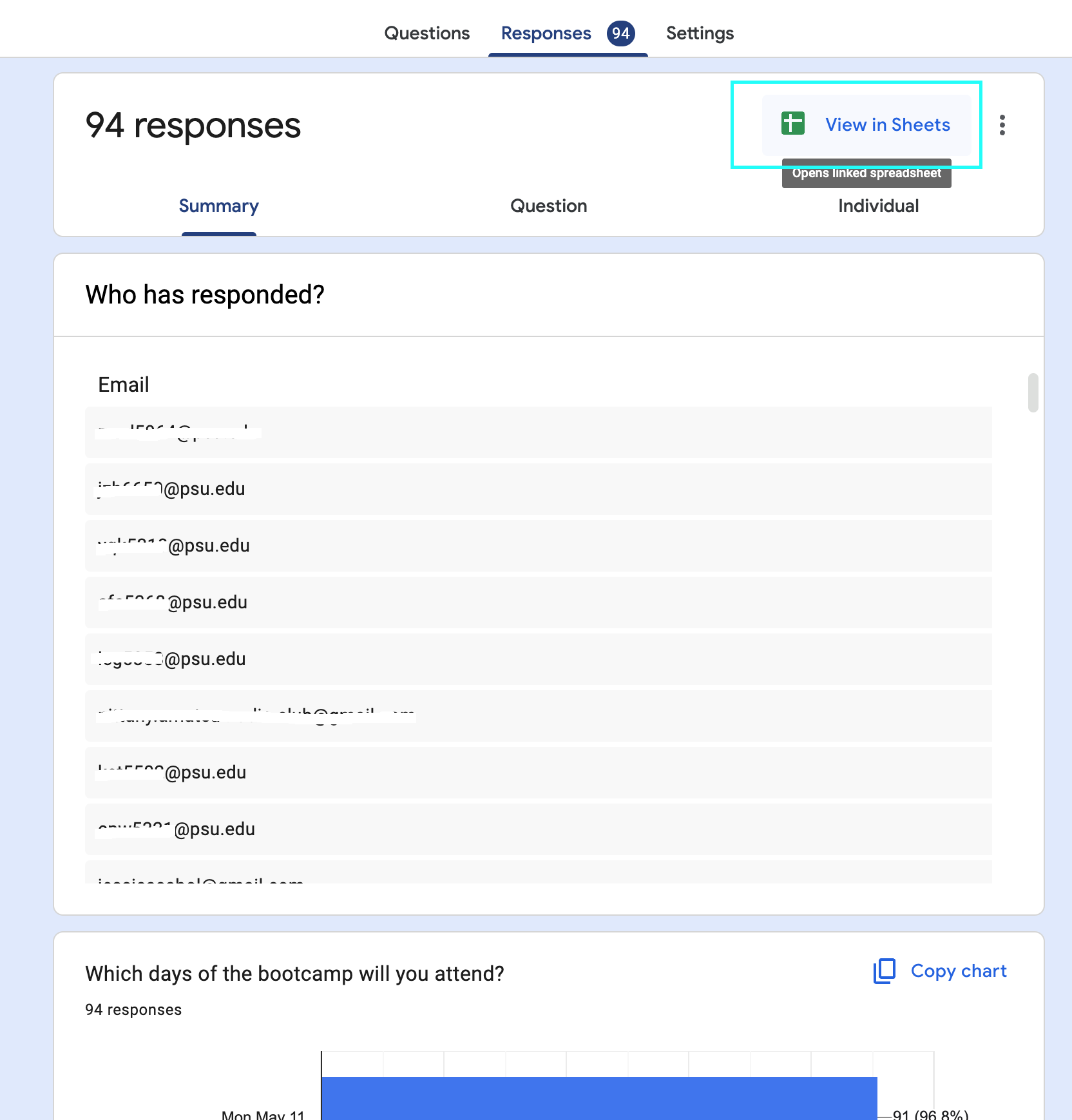
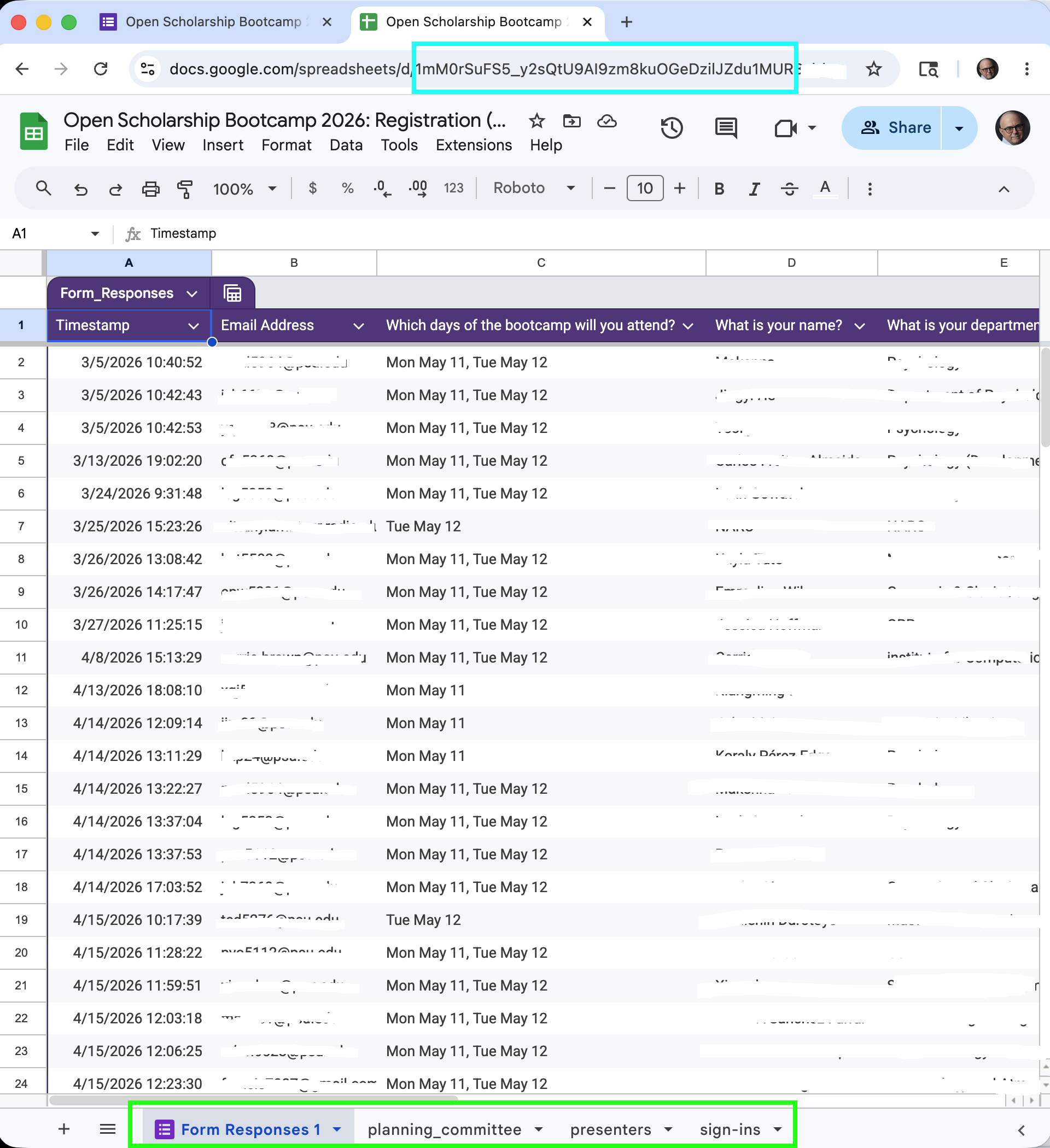
Gathering: Google Forms
Gathering: Google Forms
Gathering: Google Forms
Gathering: Google Forms
- Recommended practice:
- Save raw data locally in
data/raw_data - Save as tab- (
.tsv) or comma-separated (.csv) ASCII or utf8 text format - Use snake-case (
snake_case.csv) or hyphen-separated file names- NO SPACES OR SPECIAL CHARACTERS
- Save raw data locally in
Under the hood
- Gathering & cleaning
- Visualizing
- Gathering from Qualtrics
Your turn
Project-level organization
Tips & tricks
- Divide & conquer
- Individual files
- Small code chunks
- Tell the story (be kind to your future, forgetful, self)
- Functions!
Tips & tricks
- Use cross-referencing
- Quarto can link to figures you’ve already generated, like Figure 12.
- Great for HTML, but also works for Word & LaTex/PDF docs.
- Render; fix bugs; save.
- Automate reference generation
FAQs
- Do I have to use git or GitHub?
- No
- Especially not for identifiable data2
- But free & easy web hosting is attractive
- Can I use Quarto alongside…
- Python via Jupyter notebooks: Yes
- Shell scripts: Yes
- LaTex: Yes
FAQs
Considerations
Considerations
Wrap up
Principles
- Don’t fool yourself!
- DRY WIT
- Automate
- Show your work
- Divide & conquer (bite-sized chunks)
- Increase your bus number
Toward more reproducible scholarly workflows
Quarto (Part II): Making useful things reproducibly
Rick Gilmore
rog1 AT-SYMBOL psu PERIOD edu
114 Moore
github.com/gilmore-lab
github.com/psu-psychology
github.com/penn-state-open-science
Resources
About
This talk was produced using Quarto version 1.8.27, using the RStudio Integrated Development Environment (IDE), version 2026.4.0.526.
The source files are in R and R Markdown, then rendered to HTML using the revealJS framework. The HTML slides are hosted in a GitHub repo and served by GitHub pages: https://penn-state-open-science.github.io/bootcamp-2026-quarto-II/
Packages
We used R v. 4.6.0 (R Core Team, 2026) and the following R packages: gt v. 1.3.0 (Iannone et al., 2026), kableExtra v. 1.4.0 (Zhu, 2024), qrcode v. 0.3.0 (Onkelinx & Teh, 2024), qualtRics v. 3.2.2 (Ginn, O’Brien, & Silge, 2025), renv v. 1.2.2 (Ushey & Wickham, 2026), rmarkdown v. 2.31 (Allaire et al., 2026; Xie, Allaire, & Grolemund, 2018; Xie, Dervieux, & Riederer, 2020), tidyverse v. 2.0.0 (Wickham et al., 2019).
References
Footnotes
GitHub pages is free for public projects.
Add private files to
.gitignorePython & Jupyter notebooks workshop Today @ 3 pm: details
Alaina Pearce has implemented this.

Open Scholarship Bootcamp 2026 • Day 2 | © 2026 Rick Gilmore under CC BY 4.0